The development of biocatalytic ways to alter cheap proteogenic amino acids into bioactive synthons containing amide and carboxyl groups, which might be exploited for pharmaceutical, agrochemical, and other industrial uses, has sparked an increased scholarly interest. Many widely used synthetic and abiotic processes might benefit from enzymes that can directly catalyze the production of amide or carboxyl groups. This article looks at recent research published in ACS Catalysis.

Image Credit: Sergey Tarasov/Shutterstock.com
Once engineered into microbes, these might lead to the continuous and sustainable synthesis of diverse compounds and functions, such as α-keto and α-hydroxyl acids, stereoisomers of natural and non-natural amino acids, alcohols, peptides, amines, and so on.
Aminolysis of esters in organic solvents employing lipases is the most common enzymatic approach to amide bond production. In aqueous conditions, carboxylic acid reductases (CARs) may biocatalytically convert carboxylic acid to amides by forming reactive acyl intermediates with costly cofactors such adenosine triphosphate (ATP) and nicotinamide adenine dinucleotide phosphate (NADH).
Figure 1 shows a synthetically challenging one-pot reaction with a bio-chemo-bio catalytic route that was reported to address the lack of biosynthetic protocols for indolic amides and carboxylic acids.
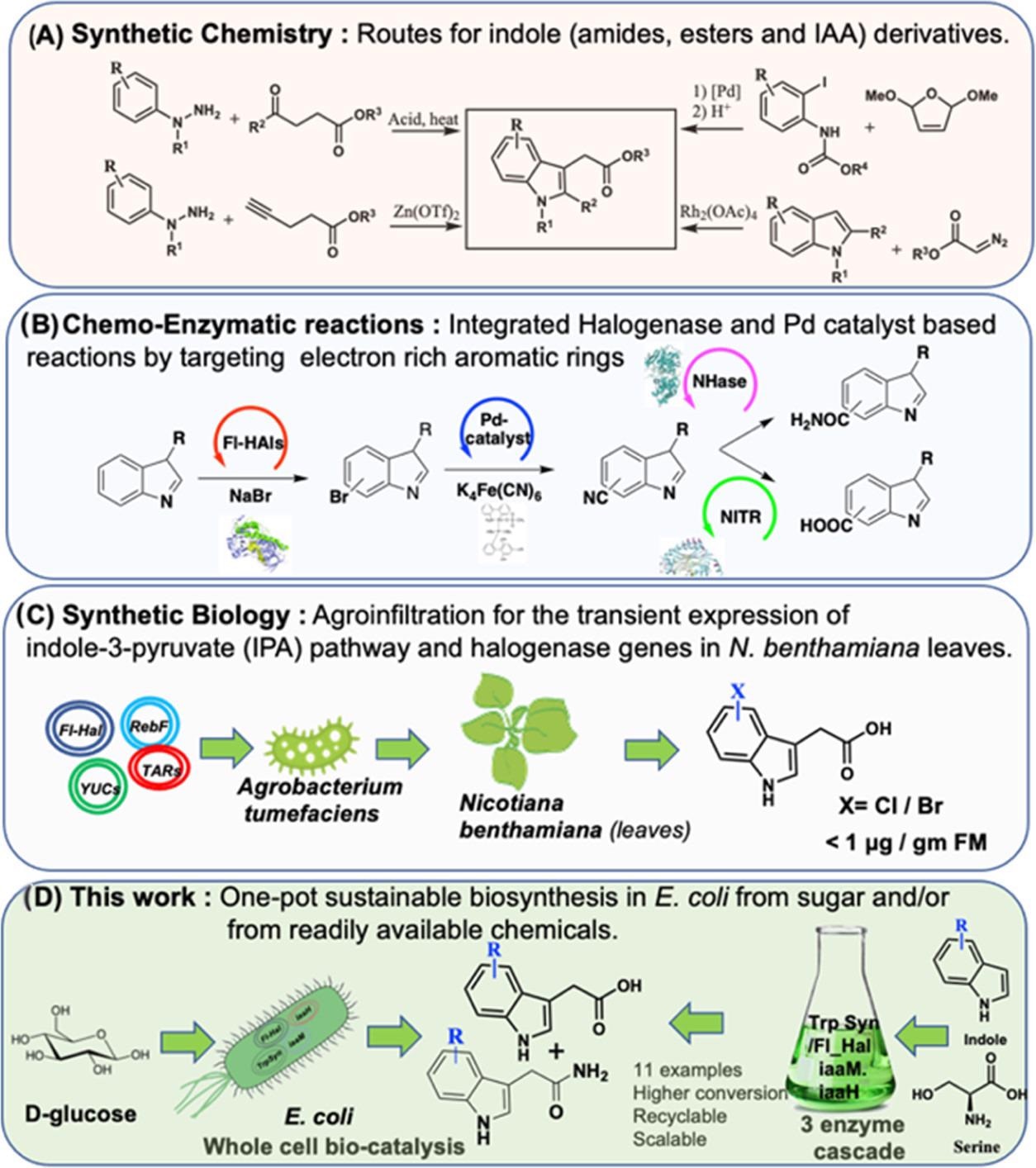
Figure 1. Chemical and biological approaches to IAA. (A) Chemical synthesis via Fisher indole synthesis and Heck or Buchwald modifications. (B) Chemo-biocatalytic routes to indole amides and carboxylic acids via halogenase-based bromination followed by cyanation with Pd catalysts. The cyano groups are converted to amide or carboxyl acid via nitrile hydratase (NHase) or nitrilase (NITR). (C) Metabolic engineering of IAA pathway genes in a plant-based compatible host organism. (D) First report of an efficient de novo biosynthesis of IAA and its derivative via the IAM pathway in engineered E. coli. Three-enzyme one-pot biotransformation approaches to IAA derivatives for large-scale synthesis are also shown. Image Credit: Menon, et al., 2022
This research shows how to recreate and use difficult plant-based natural product pathways in industrial hosts to turn on innovative chemistries for large-scale synthesis.
For example, tryptophan-2-monooxygenase (iaaM) acts on l-tryptophan to produce indole-3-acetamide (IAM), which is then converted to indole-3-acetic acid (IAA) by indole-3-acetamide hydrolase (iaaH) depicts in Figure 2.
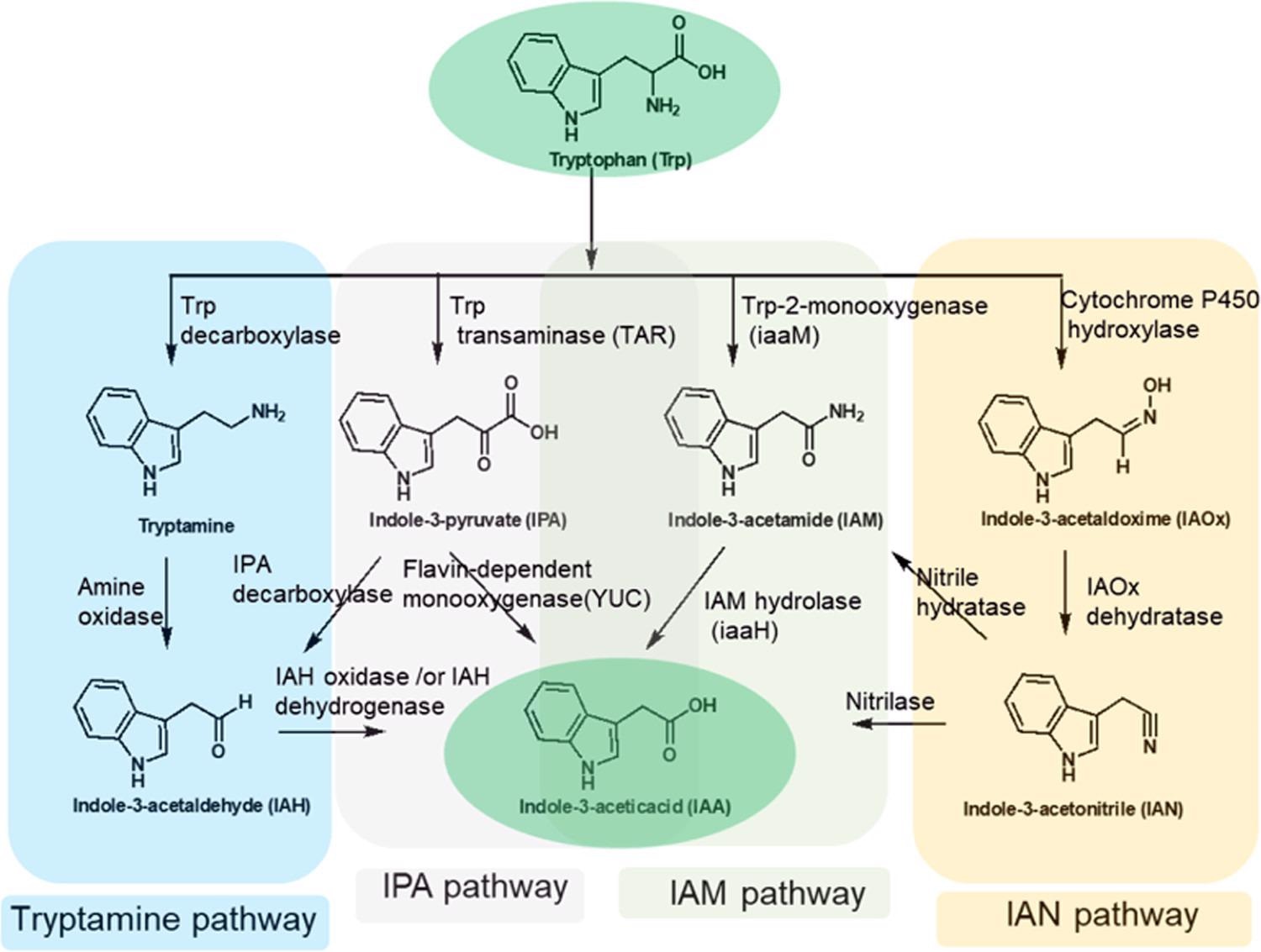
Figure 2. Tryptophan-dependent biosynthetic pathways toward indole-3-acetic acid in plants and plant-associated microbiomes. IAM and IPA pathways are the major route(s) to IAA formation in plants and plant-associated microorganisms. Image Credit: Menon, et al., 2022
Results
From the plant and microbial genome database repository, researchers identified iaaM and iaaH candidate genes from plants and plant-associated species of bacteria.
In vitro tests using tryptophan and tyrosine were used to investigate the kinetic characteristics of the purified SUMO-iaaM fusion protein. Table 1 shows that SUMO-iaaM has an apparent kcat of 8.4 s-1 for tryptophan and 0.3 ± 0.05 s–1 for tyrosine.
Table 1. Kinetic Parameters Determined for SUMO-iaaM, SUMO-iaaH Wild-Type Enzymes, and Their Important Mutants (Errors Are S.E.M., Where n = 3). Source: Menon, et al., 2022
| enzyme |
substrate |
kcat (s–1) |
KM (µM) |
kcat/KM (mM–1 s–1) |
| WT SUMO-iaaM |
tryptophan |
>8.4 (apparent kcat) |
42.8 ± 0.3 |
196.3 ± 4.9 (estimated) |
| tyrosine |
0.3 ± 0.05 |
112.2 ± 0.4 |
2.7 ± 0.4 |
| R98K-iaaM |
tryptophan |
0.15 ± 0.03 |
140.8 ± 1.8 |
1.06 ± 0.2 |
| F244A-iaaM |
tryptophan |
0.09 ± 0.01 |
>200 |
ND |
| L478E-iaaM |
tryptophan |
0.08 ± 0.01 |
>200 |
ND |
| enzyme |
substrate |
kcat (min–1) |
KM (µM) |
kcat/KM (mM–1 min–1) |
| WT SUMO-iaaH |
IAM |
2.8 ± 0.4 |
<25 |
111.1 ± 16.2 (estimated) |
| benzamide |
0.15 ± 0.02 |
61.5 ± 0.5 |
2.4 ± 0.3 |
| S151T-iaaH |
IAM |
0. 56 ± 0.08 |
135.2 ± 1.3 |
4.1 ± 0.6 |
| S152A-iaaH |
IAM |
0.23 ± 0.06 |
142.2 ± 0.9 |
1.6 ± 0.4 |
Following that, the SUMO fusion proteins were purified to homogeneity, enabling additional kinetic and mutagenesis analyses of these enzymes.
Initially, alanine-substituted mutant enzymes for Arg98, Phe244, Lys365, Trp466, Phe476, Leu478, and Trp519 residues were produced, and their activity was evaluated against tryptophan and tyrosine to determine the relevance of these residues in substrate binding and recognition.
Alanine substitutions at Leu478 and Trp519, on the other hand, were shown to have less influence on the iaaM enzyme's relative activity, indicating that these residues play a minimal part in catalysis.
The essential active site residues in iaaM include Arg98, Phe244, Lys365, Trp466, Phe476, Leu478, and Trp519, which are situated within a 5 Å distance between the substrate and the FAD-binding pocket, as shown in Figures 3.
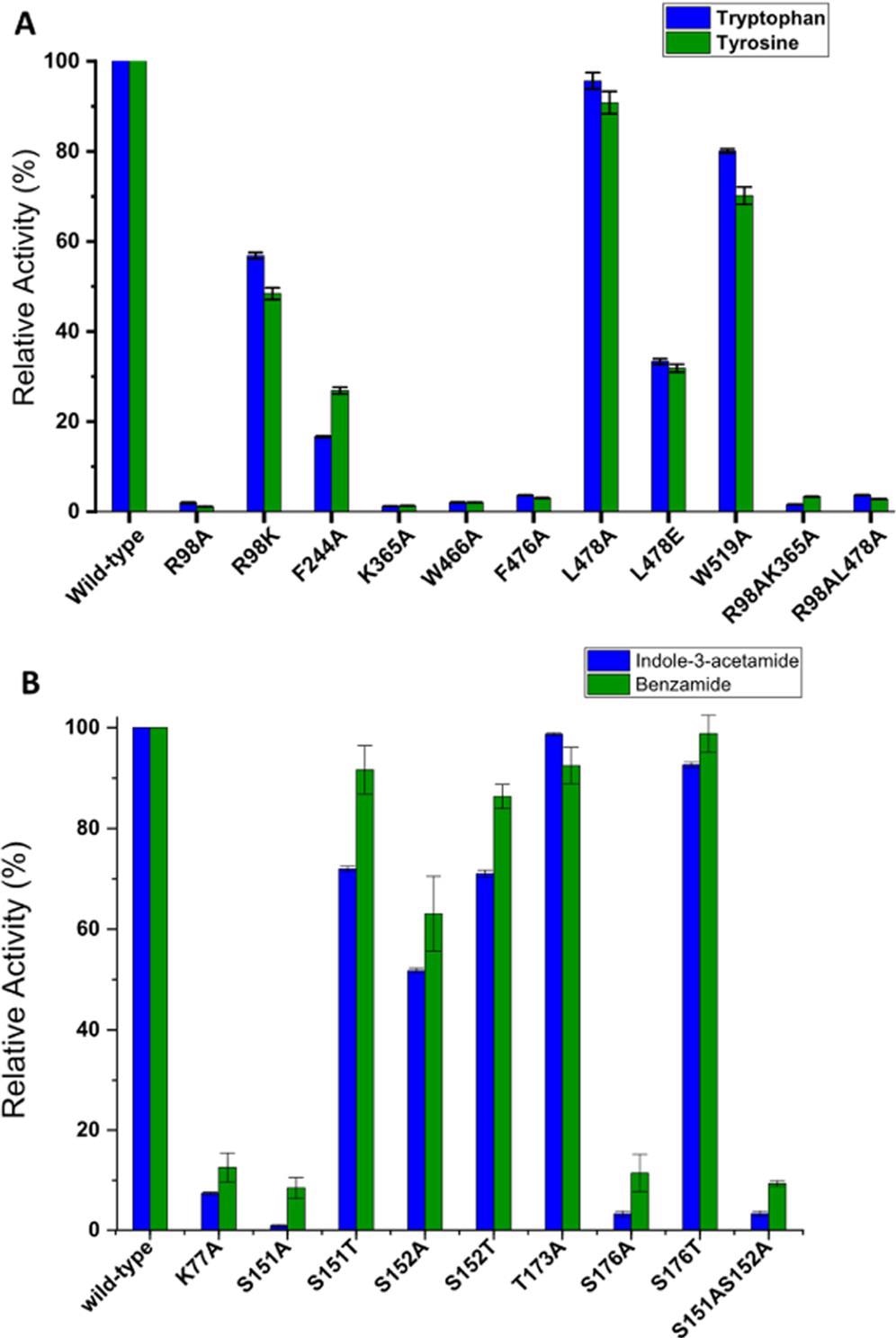
Figure 3. Relative activity of tryptophan-2-monooxygenase (iaaM) (A) and indole-3-acetamide hydrolase (iaaH) (B) mutant enzymes in comparison to wild-type in the presence of different substrates. The substrate conversion of wild-type enzyme is taken as 100% in all cases. The reaction was performed using purified enzymes (10 μm) by incubating them with 1 mM substrate for 30 min at 25 °C with gentle agitation (180 rpm). The total volume of reaction was 500 μL, which was prepared in a 50 mM Tris-HCl buffer solution at pH 7.8. The reaction was performed in duplicate with error bars indicating the standard deviation of the HPLC measurements. Image Credit: Menon, et al., 2022
Researchers performed whole-cell tests using SUMO-iaaM and SUMO-iaaH proteins in E. coli cells to further investigate the scope of establishing an in vivo reaction with active IAM pathway enzymes are illustrated in Figure 4.
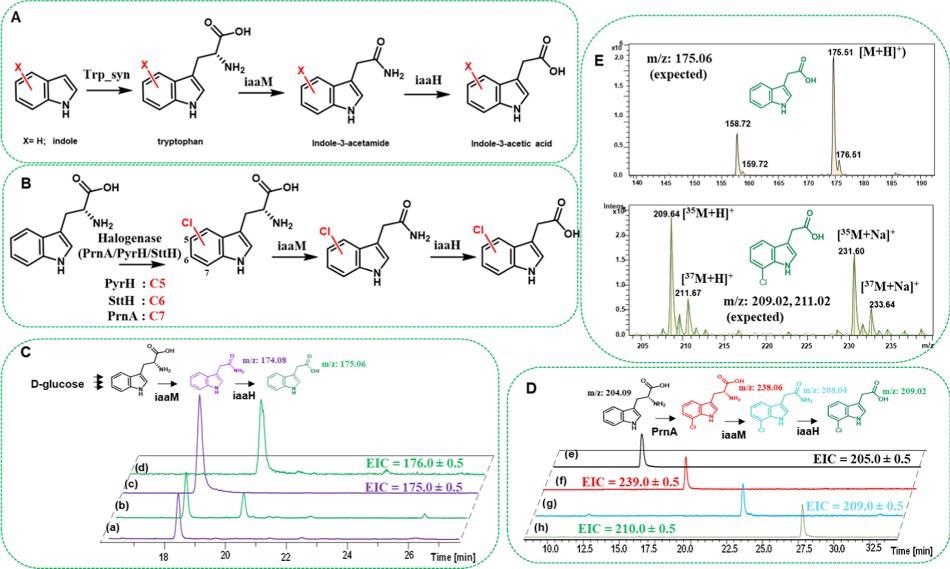
Figure 4. Tryptophan synthase (Trp_syn)- and flavin-dependent halogenase (Fl-Hal)-based pathways (A and B, respectively) exploited in this study for amide and carboxylic acid bond formation in E. coli cells via one-pot cascade reactions. (C) HPLC traces showing in vivo formation of indole-3-acetamide and indole-3-acetic acid in E. coli cell cultures overexpressed with SUMO-iaaM or SUMO-iaaM along with SUMO-iaaH enzymes (trace a is iaaM and trace b is iaaM/iaaH duet expression). Traces c and d are the extracted ion chromatograms for indole-3-acetamide and indole-3-acetic acid, respectively. (D) Extracted ion chromatograms showing in vivo production of 7-chlorotryptophan, 7-chloro indole-3-acetamide, and 7-chloro indole-3-acetic acid after feeding cells with tryptophan. Traces e and f are the extracted ion chromatograms for tryptophan and 7-chlorotryptophan from the cell cultures that carry PrnA Fl-Hal plasmid only, respectively. Trace g is the extracted ion chromatogram for 7-chloro indole-3-acetamide from the cell culture that carries PrnA and SUMO-IaaM plasmids, whereas h is that of 7-chloro indole-3-acetic acid from the cultures that carry PrnA and SUMO-IaaM/SUMO-IaaH duet plasmids. For visual clarity, in (C, D), molecules are drawn with similar colors to the HPLC traces they represent. (E) ESI-MS spectra for indole-3-acetic acid and 7-chloro indole-3-acetic acid produced via cell cultures. The experimental protocols are detailed in the Materials and Methods section. Image Credit: Menon, et al., 2022
The initial strategy of co-expressing Trp syn with the pET-SUMO-iaaM and pCDF-SUMO-iaaH constructs appeared to improve IAM and IAA titers. However, the best results were obtained when the Trp syn-overexpressed cell supernatant was used for whole-cell biosynthesis, as shown in Table 2.
Table 2. Reconstructed IAM Pathway and HPLC Conversion Values for the Formation of Chlorinated and Nonchlorinated Indole-3-Acetamide and Indole-3-Acetic Acid Molecules under Various Assay Conditions via In Vivo and In Vitro Cascade Reactions in E. coli cells. Source: Menon, et al., 2022
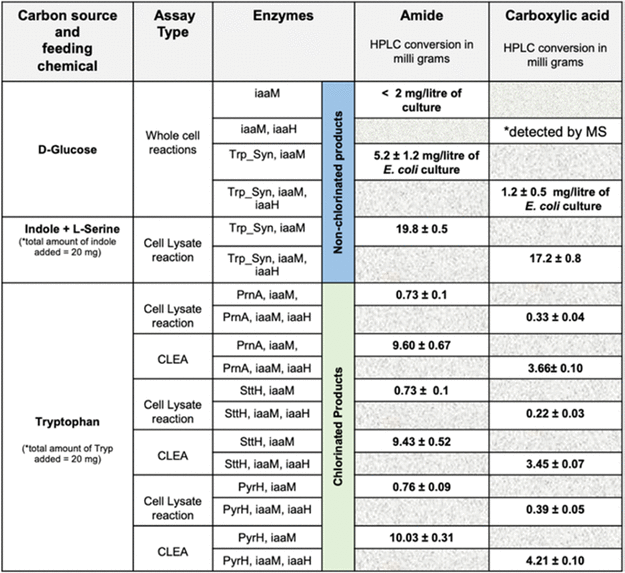
Trp syn accepted chloro- and bromosubstituted indoles with the halogen atom at positions 7, 6, 5, and 4 to form substituted tryptophan compounds. These compounds were converted to IAM and IAH derivatives by the downstream activity of iaaM and IaaH enzymes (Table 3).
Table 3. Three-Enzyme One-Pot Cell Lysate Cascade Reaction with Tryptophan Synthase (Trp_syn), Tryptophan-2-Monooxygenase (iaaM), and Indole-3-Acetamide Hydrolase (iaaH) by Feeding Substituted Indole and Serine Molecules. Source: Menon, et al., 2022
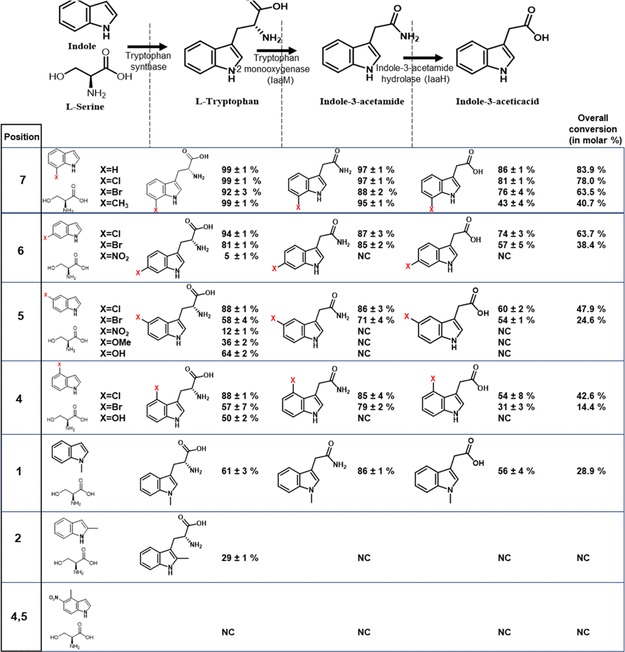
The addition of CLEA increased PrnA’s 7-chlorotryptophan conversion to 61%, compared to the 9% conversion seen in the cell lysate tests, are presented in Table 2 and Figure 5.
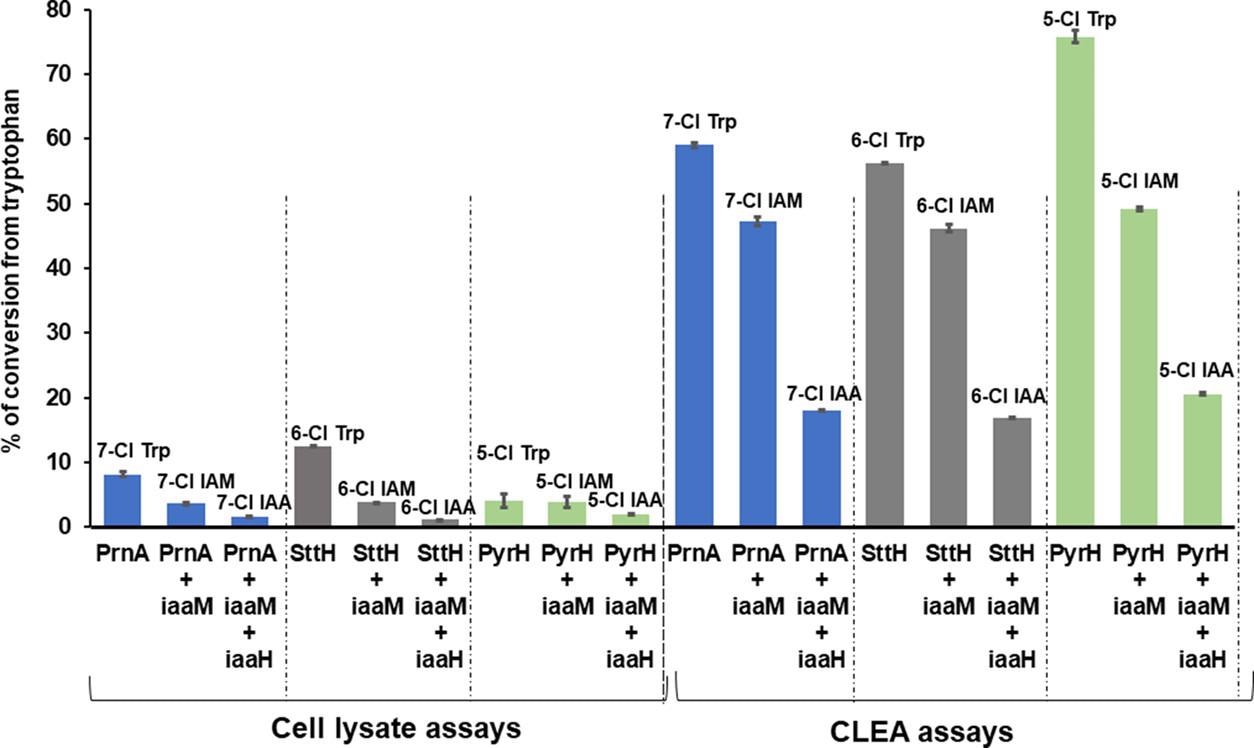
Figure 5. Comparison of HPLC conversion for three-enzyme cascade reactions with Fl-Hal (PrnA, PyrH, or SttH) and IAM pathway enzyme (iaaM and/or iaaH) combinations using either cell lysate reactions or CLEA catalysts. Reactions were performed by feeding 20 mg of tryptophan in 100 mM KPi buffer at pH 7.8 for 4 days at 30 °C. The experimental protocols are detailed in the Materials and Methods section. Image Credit: Menon, et al., 2022
Conclusion
Researchers have recreated and established a plant-associated biosynthetic pathway in E. coli that generates numerous substituted IAM and IAA molecules that were previously unavailable.
Researchers have opened future avenues to engineer these enzymes to produce many other value-added molecules from different amino acid precursors, in addition to identifying an appropriate solution to create a highly overexpressed and soluble E. coli version for IAM pathway enzymes and performing detailed mutagenesis analysis. To adapt the enzyme to accept natural and non-natural amino acids, scientists are now using site-directed and random mutagenesis techniques.
Researchers have shown multiple methods to convert readily accessible compounds such as indole and tryptophan to synthesis numerous IAM and IAA analogs by combining the process with tryptophan synthase and flavin-dependent regiospecific halogenases.
Scientists further demonstrated that these enzymes are compatible with in vivo reactions. The cell lysate and CLEA-based techniques broadened the present spectrum of IAM pathway enzymes.
The CLEA-based cascade systems established in this study are very adaptable, scalable, measurable, and reusable for large-scale synthesis and commercial applications, which will stimulate the attention of the biocatalysis community and plant biotechnology researchers to examine other flavin-dependent enzymes and pathways found in plants.
Journal Reference:
Menon, N., Richmond, D., Rahman, M. R. and Menon, B. R (2022) Versatile and Facile One-Pot Biosynthesis for Amides and Carboxylic Acids in E. coli by Engineering Auxin Pathways of Plant Microbiomes. ACS Catalysis, 12, pp.2309-2319. Available Online: https://pubs.acs.org/doi/10.1021/acscatal.1c04901.
References and Further Reading
- Craven, E. J., et al. (2021) Programmable late-stage C–H bond functionalization enabled by integration of enzymes with chemocatalysis. Nature Catalysis, 4, pp.385–394. doi.org/10.1038/s41929-021-00603-3.
- Bartsch, S & Bornscheuer, U T (2010) Mutational analysis of phenylalanine ammonia lyase to improve reactions rates for various substrates. Protein Engineering, Design and Selection, 23, pp. 929– 933. doi.org/10.1093/protein/gzq089.
- Weise, N. J., et al. (2017) Turner, N. J. Zymophore identification enables the discovery of novel phenylalanine ammonia lyase enzymes. Scientific Reports, 7, p.13691. doi.org/10.1038/s41598-017-13990-0.
- Cheng, Z., et al. (2020) Recent Advances and Promises in Nitrile Hydratase: From Mechanism to Industrial Applications. Frontiers in Bioengineering & Biotechnology, 8, p. 352. doi.org/10.3389/fbioe.2020.00352.
- Intasian, P., et al. (2021) Enzymes, In Vivo Biocatalysis, and Metabolic Engineering for Enabling a Circular Economy and Sustainability. Chemical Reviews, 121, pp. 10367–10451. doi.org/10.1021/acs.chemrev.1c00121.
- Song, W., et al. (2020) Biocatalytic derivatization of proteinogenic amino acids for fine chemicals. Biotechnology Advances, 40, p. 107496. doi.org/10.1016/j.biotechadv.2019.107496.
- Parmeggiani, F., et al. (2015) Synthesis of D- and L-phenylalanine derivatives by phenylalanine ammonia lyases: a multienzymatic cascade process. Angewandte Chemie International Edition in English, 54, pp. 4608–4611. doi.org/10.1002/anie.201410670.
- Boville, C. E., et al. (2018) Engineered Biosynthesis of beta-Alkyl Tryptophan Analogues. Angewandte Chemie International Edition in English, 57, pp. 14764–14768. doi.org/10.1002/anie.201807998.
- Liljeblad, A., et al. (2010) Formation and hydrolysis of amide bonds by lipase A from Candida antarctica; exceptional features. Organic and Biomolecular Chemistry, 8, pp. 886–895. doi.org/10.1039/b920939p.
- Zeng, S., et al. (2018) Amide Synthesis via Aminolysis of Ester or Acid with an Intracellular Lipase. ACS Catalysis, 8, pp. 8856–8865. doi.org/10.1021/acscatal.8b02713.
- Wood, A. J. L., et al. (2017) Adenylation Activity of Carboxylic Acid Reductases Enables the Synthesis of Amides. Angewandte Chemie International Edition in English, 56, pp. 14498–14501. doi.org/10.1002/anie.201707918.
- Ramsden, J. I., et al. (2019) Biocatalytic N-Alkylation of Amines Using Either Primary Alcohols or Carboxylic Acids via Reductive Aminase Cascades. Journal of the American Chemical Society, 141, pp. 1201–1206. doi.org/10.1021/jacs.8b11561.
- Finnigan, W., et al. (2017) N. J. Characterization of Carboxylic Acid Reductases as Enzymes in the Toolbox for Synthetic Chemistry. ChemCatChem, 9, pp. 1005–1017. doi.org/10.1002/cctc.201601249.
- Pongpamorn, P., et al. (2021) Carboxylic Acid Reductase Can Catalyze Ester Synthesis in Aqueous Environments. Angewandte Chemie International Edition in English, 60, pp.5749–5753. doi.org/10.1002/anie.202013962.
- Petchey, M. R., et al. (2020) Biocatalytic Synthesis of Moclobemide Using the Amide Bond Synthetase McbA Coupled with an ATP Recycling System. ACS Catalysis, 10, pp. 4659–4663. doi.org/10.1021/acscatal.0c00929.
- Petchey, M., et al. (2018) The Broad Aryl Acid Specificity of the Amide Bond Synthetase McbA Suggests Potential for the Biocatalytic Synthesis of Amides. Angewandte Chemie International Edition in English, 57, pp.11584–11588. doi.org/10.1002/anie.201804592.
- Philpott, H. K., et al. (2018) A versatile biosynthetic approach to amide bond formation. Green Chemistry, 20, pp. 3426–3431. doi.org/10.1039/C8GC01697F.
- Tsuji, J., et al. (1965) Organic Syntheses by Means of Noble Metal Compounds. XII.1 Reaction of the Cyclooctadiene-Palladium Chloride Complex with Ethyl Malonate. Journal of the American Chemical Society, 87, pp. 3275–3276. doi.org/10.1021/ja01092a067.
- Trost, B. M & Fullerton, T J (1973) New synthetic reactions. Allylic alkylation. Journal of the American Chemical Society, 95, pp. 292–294. doi.org/10.1021/ja00782a080.
- Chen, D., et al. (2018) One-Pot Synthesis of Indole-3-acetic Acid Derivatives through the Cascade Tsuji-Trost Reaction and Heck Coupling. The Journal of Organic Chemistry, 83, pp. 6805–6814. doi.org/10.1021/acs.joc.8b01056.
- Davis, K., et al. (2020) Nicotiana benthamiana as a Transient Expression Host to Produce Auxin Analogs. Frontiers in Plant Science, 11, p. 581675. doi.org/10.3389/fpls.2020.581675.
- Lehmann, T., et al. (2010) Indole-3-acetamide-dependent auxin biosynthesis: a widely distributed way of indole-3-acetic acid production? European Journal of Cell Biology, 89, pp. 895–905. doi.org/10.1016/j.ejcb.2010.06.021.
- De Rybel, B., et al. (2009) The past, present, and future of chemical biology in auxin research. ACS Chemical Biology, 4, pp. 987–998. doi.org/10.1021/cb9001624.
- Duca, D., et al. (2014) Indole-3-acetic acid in plant-microbe interactions. Antonie van Leeuwenhoek, 106, pp. 85–125. doi.org/10.1007/s10482-013-0095-y.
- Liu, Y. Y., et al. (2016) Variation in Indole-3-Acetic Acid Production by Wild Saccharomyces cerevisiae and S. paradoxus Strains from Diverse Ecological Sources and Its Effect on Growth. PLoS One, 11, p. e0160524. doi.org/10.1371/journal.pone.0160524.
- Fu, S., et al. (2015) Indole-3-acetic acid: A widespread physiological code in interactions of fungi with other organisms. Plant Signaling & Behavior, 10, p. e1048052. doi.org/10.1080/15592324.2015.1048052.
- Lewis, J H & Stine, J G (2013) Nonsteroidal Antiinflammatory Drugs and Leukotriene Receptor Antagonists. In Drug-Induced Liver Disease, 3rd ed.; Kaplowitz, N.; DeLeve, L. D., Eds.; Academic Press: Boston; Chapter 22, pp 369–401.
- Bunsangiam, S., et al. (2019) Biosynthetic Pathway of Indole-3-Acetic Acid in Basidiomycetous Yeast Rhodosporidiobolus fluvialis. Mycobiology, 47, pp. 292–300. doi.org/10.1080/12298093.2019.1638672.
- Myo, E. M., et al. (2019) Indole-3-acetic acid production by Streptomyces fradiae NKZ-259 and its formulation to enhance plant growth. BMC Microbiology, 19, p. 155. doi.org/10.1186/s12866-019-1528-1.
- Shao, J., et al. (2015) Analysis and cloning of the synthetic pathway of the phytohormone indole-3-acetic acid in the plant-beneficial Bacillus amyloliquefaciens SQR9. Microbial Cell Factories, 14, p. 130. doi.org/10.1186/s12934-015-0323-4.
- Romasi, E F & Lee, J (2013) Development of indole-3-acetic acid-producing Escherichia coli by functional expression of IpdC, AspC, and Iad1. JMB Journal of Microbiology and Biotechnology, 23, pp.1726–1736. doi.org/10.4014/jmb.1308.08082.
- Guo, D., et al. (2019) De Novo Biosynthesis of Indole-3-acetic Acid in Engineered Escherichia coli. Journal of Agricultural and Food Chemistry, 67, pp. 8186–8190, doi.org/10.1021/acs.jafc.9b02048.
- Mashiguchi, K., et al. (2011) The main auxin biosynthesis pathway in Arabidopsis. Proceedings of the National Academy of Sciences of the United States of America, 108, pp. 18512–18517. doi.org/10.1073/pnas.1108434108.
- Imada, E. L., et al. (2017) Indole-3-acetic acid production via the indole-3-pyruvate pathway by plant growth promoter Rhizobium tropici CIAT 899 is strongly inhibited by ammonium. Research in Microbiology, 168, pp. 283–292. doi.org/10.1016/j.resmic.2016.10.010.
- Takahashi, Y., et al. (2010) Characterization of five polyamine oxidase isoforms in Arabidopsis thaliana. Plant Cell Reports, 29, pp. 955–965. doi.org/10.1007/s00299-010-0881-1.
- Raut, V., et al. (2017) Plant growth promotion using microbial IAA producers in conjunction with azolla: a novel approach. Chemical and Biological Technologies in Agriculture, 4, p. 1. doi.org/10.1186/s40538-016-0083-3.
- Gaweska, H. M., et al. (2013) Structure of the flavoprotein tryptophan 2-monooxygenase, a key enzyme in the formation of galls in plants. Biochemistry, 52, pp. 2620–2626, doi.org/10.1021/bi4001563.
- Emanuele, J. J., et al. (1995) Purification and characterization of the flavoprotein tryptophan 2-monooxygenase expressed at high levels in Escherichia coli. Archives of Biochemistry and Biophysics, 316, pp. 241–248. doi.org/10.1006/abbi.1995.1034.
- Mashiguchi, K., et al. (2019) Agrobacterium tumefaciens Enhances Biosynthesis of Two Distinct Auxins in the Formation of Crown Galls. Plant Cell Physiology, 60, pp. 29–37. doi.org/10.1093/pcp/pcy182.
- Hutcheson, S. W & Kosuge, T (1985) Regulation of 3-indoleacetic acid production in Pseudomonas syringae pv. savastanoi. Purification and properties of tryptophan 2-monooxygenase. Journal of Biological Chemistry, 260, pp. 6281–6287. doi.org/10.1016/S0021-9258(18)88968-2.
- Emanuele, J J & Fitzpatrick, P F (1995) Mechanistic studies of the flavoprotein tryptophan 2-monooxygenase. 2. pH and kinetic isotope effects. Biochemistry, 34, pp. 3716–3723. doi.org/10.1021/bi00011a029.
- Sobrado, P & Fitzpatrick, P F (2003) Analysis of the role of the active site residue Arg98 in the flavoprotein tryptophan 2-monooxygenase, a member of the L-amino oxidase family. Biochemistry, 42, pp. 13826–13832. doi.org/10.1021/bi035299n.
- Niu, H., et al. (2019) Metabolic engineering for improving L-tryptophan production in Escherichia coli. Journal of Industrial Microbiology and Biotechnology, 46, pp. 55–65. doi.org/10.1007/s10295-018-2106-5.
- Schoppel, K., et al. (2021) Metabolic control analysis of L-tryptophan producing Escherichia coli applying targeted perturbation with shikimate. Bioprocess and Biosystems Engineering, 44, pp. 2591–2613. doi.org/10.1007/s00449-021-02630-7.
- Dick, M., et al. (2019) Tailoring Tryptophan Synthase TrpB for Selective Quaternary Carbon Bond Formation. Journal of the American Chemical Society, 141, pp. 19817–19822. doi.org/10.1021/jacs.9b09864.
- Watkins-Dulaney, E., (2021) Tryptophan Synthase: Biocatalyst Extraordinaire. ChemBioChem, 22, pp. 5–16. doi.org/10.1002/cbic.202000379.
- Francis, D., et al. (2017) An Engineered Tryptophan Synthase Opens New Enzymatic Pathways to beta-Methyltryptophan and Derivatives. ChemBioChem, 18, pp. 382–386. doi.org/10.1002/cbic.201600471.
- Latham, J., et al. (2016) Integrated catalysis opens new arylation pathways via regiodivergent enzymatic C-H activation. Nature Communications, 7, p. 11873. doi.org/10.1038/ncomms11873.
- Sharma, S. V., et al. (2017) Living GenoChemetics by hyphenating synthetic biology and synthetic chemistry in vivo. Nature Communications, 8, p. 229. doi.org/10.1038/s41467-017-00194-3.
- Menon, B. R. K., et al. (2020) Halogenases for biosynthetic pathway engineering: Toward new routes to naturals and non-naturals. Catalysis Reviews, pp. 1–59. doi.org/10.1080/01614940.2020.1823788.
- Mondal, D., et al. (2021) Flavin-dependent halogenases catalyze enantioselective olefin halocyclization. Nature Communications, 12, p. 3268. doi.org/10.1038/s41467-021-23503-3.
- Shepherd, S. A., et al. (2015) Extending the biocatalytic scope of regiocomplementary flavin-dependent halogenase enzymes. Chemical Science, 6, pp. 3454–3460. http://doi.org/10.1039/c5sc00913h
- Schnepel, C & Sewald, N (2017) Enzymatic Halogenation: A Timely Strategy for Regioselective C-H Activation. Chemistry—A European Journal, 23, pp. 12064–12086. doi.org/10.1002/chem.201701209.
- Shepherd, S. A., et al. (2016) A Structure-Guided Switch in the Regioselectivity of a Tryptophan Halogenase. ChemBioChem, 17, pp. 821–824. doi.org/10.1002/cbic.201600051.
- Gkotsi, D. S., et al. (2019) A marine viral halogenase that iodinates diverse substrates. Nature Chemistry, 11, pp. 1091–1097. doi.org/10.1038/s41557-019-0349-z.
- Brown, S & O’Connor, S. E (2015) Halogenase Engineering for the Generation of New Natural Product Analogues. ChemBioChem, 16, pp. 2129–2135. doi.org/10.1002/cbic.201500338.